Is anti-viral interferon response a boon to SARS-CoV-2?
Viruses are infamous for using our own system against us. Most viruses do that, however individual viruses have their own strategies in how they will achieve it. The story for any virus begins with attaching to a receptor on our cells and then gaining entry. Once they enter they multiply and make proteins that are structurally and functionally important for the virus. They assemble into new virions and exit the cells to infect nearby cells or shed from our body to a new host.
Illustrated by @scienceblocks using following images -
Crime by Aby Badali | CC BY-SA 2.5
SARS-CoV-2 by CDC/ Alissa Eckert, MS; Dan Higgins, MAM | Public Domain. Enterocyte by Boumphreyfr | CC BY-SA 3.0
The life cycle of SARS-CoV-2
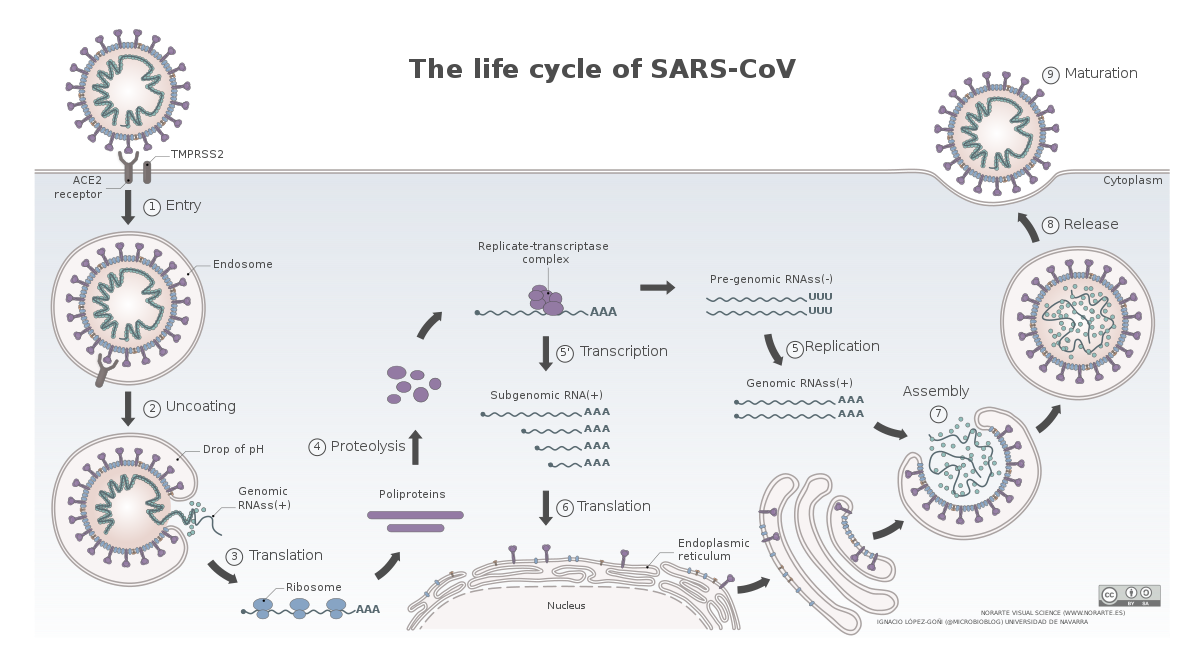
Image by Vega Asensio | CC BY-SA 4.0
The novel coronavirus, SARS-CoV-2, have a similar life cycle. Once, it enters the respiratory tract it binds to angiotensin-converting enzyme 2 (ACE2) receptor on the surface of some cells (Hoffman et al., 2020). This is followed by cleavage of spike protein on the surface of the virus via a protease TMPRSS2. This exposes a domain of spike which promotes membrane fusion between the virus and the cell. The virus hence gains entry into the cells and gets on with its business. However, which cells in the respiratory tract and other organs known to be infected by this virus express ACE2 and how is the expression of ACE2 regulated in these cells is not fully understood.
Check out this video for detailed explanation.
The paper by Zeigler et al., 2020, that we will discuss today gives us a hint about exactly that. It talks about which cells where express ACE2 and TMPRSS2, and more importantly how the expression of ACE2 might be regulated. It also gives us a hint that how the SARS-CoV-2 might actually be using the antiviral response of the cells to its own benefit. And if that is so what can this regulation tell us about who is more likely to be severely affected? What about using antiviral interferon as therapy? Which model of the disease we should or should not use when studying the disease or testing new cures?
Which cells express ACE2?
Zeigler et al., took a very interesting approach to figure out which cells express ACE2 and TMPRSS2. They have analysed the single cell RNA seq (scRNAseq) data from lungs, nasal passage and gut of humans and non-human primates. The cool thing about scRNAseq is that it looks at the genes expressed in individual cells and create clusters of cells, such that cells with similar overall gene expression fall into one cluster. Then using the known genes which are expressed only particular kind of cells you can identify what cell type a particular cluster correspond to. You may then ask the question that in which of these clusters your gene of interest is expressed in.
The cells in nasal passage and lungs that CoV-2 would love to infect
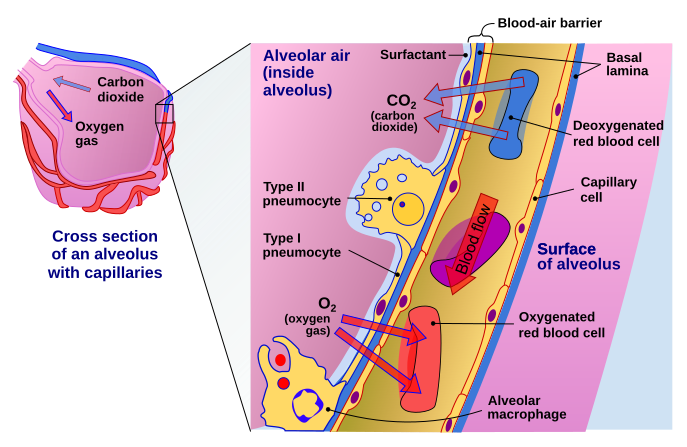
Image by Delmalani18 | [CC BY-SA 4.0]
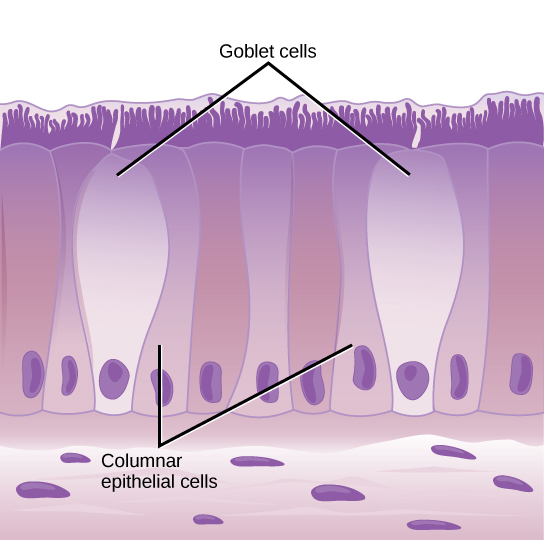
Image by CNX OpenStax | CC BY 4.0
In the lungs, the ACE2 expression was majorly found in type II pneumocytes and in ciliated cells of respiratory tract epithelium. To give you an idea of numbers about 1.4% type II pneumocytes, and 7% ciliated cells expressed ACE2. 0.8% of Type II pneumocytes and 5.3% of ciliated cells expressed both ACE2 and TMPRSS2. But that's the story of the lungs. However, if the virus has to infect it first needs to attach and infect cells close to the entry point. They identified that secretory goblet cells were the major target for SARS-CoV-2 in nasal epithelium.
The gut has target cells for CoV-2, as well
Now, if you are aware of symptoms of Covid19, the disease caused by SARS-CoV-2, you might know that it is characterised by pneumonia, fever, cough and sometimes even diarrhoea. Moreover, it has been shown that covid19 patients keep shedding the virus in stool for quite some time (Wu et al., 2020). The virus has also been shown to infect the GI tract (Xiao et al., 2020, Wang et al., 2020). Xaio et al., found that ACE2 was majorly expressed in cilia of glandular epithelium and they found the viral protein in stomach, duodenum and rectum. In line with this Zeigler et al., identified ACE2 expressing cells in the ileum of the small intestine - absorptive enterocytes.
The regulation of ACE2 by anti-viral molecules - Interferons.
ACE2 - interferon-stimulated gene?
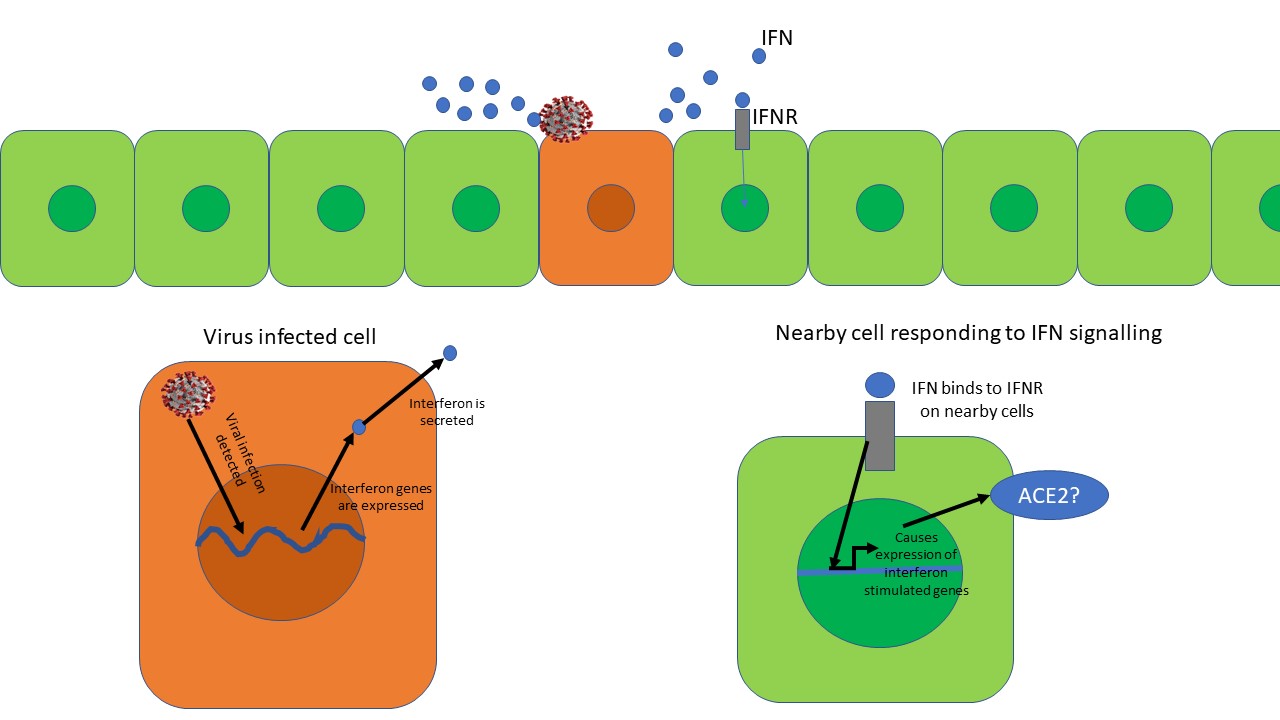
Anti-viral interferon response. The virus infected cell secretes interferon's (IFNs). The IFN binds to interferon receptor (IFNR) on nearby cells and starts expression of interferon-stimulated genes. However, is Ace2 an interferon-stimulated gene? Illustrated by @scienceblocks
There was something more interesting going on here. No matter where the authors took samples from - non-human primates (Rhesus macaque) or humans, they found that expression of ACE2 always correlated with expression of genes known to be expressed during an interferon response - a cellular defence against viral infection. Now, this is interesting, because interferons are class of molecules released by virus-infected cells to warn the nearby cells to up-regulate their viral defences. They also modulate the immune response and increase expression of MHC on cells (a molecule that displays viral peptides to T cells).
To confirm that interferon can indeed increase the expression of ACE2 in cells, Zeigler et al., treated cells derived from human nasal epithelium with interferon-gamma and interferon-alpha 2. They noticed a drastic increase in the expression of ACE2 gene in these cells. So, what does it mean if an anti-viral response of interferon can itself increase the receptor to which the virus will bind on the cells? Does it mean that the SARS-CoV-2 would use our viral defence system against itself? Is the virus being like - bring that interferon on humans! it may slow me down a bit, maybe, but it will also increase my chances to infect nearby cells? Moreover, given this information what does it mean for the interferon therapy proposed as a possible treatment against the virus? Is it really a good idea?
Viral infections increase the expression of ACE2?
That being said, what does interferon increasing expression of ACE2 mean for people with other viral infections. It is likely that people infected with any other virus may have a higher expression of ACE2 in certain cells and could be at higher risk of contracting Covid19 and becoming more severely ill? At least, in this paper, the authors show that HIV+ patients have higher expression of ACE2 in their cells. Given that HIV infection is known to cause a chronic interferon response in the body, it may explain why. Moreover, infecting non-human primates with Simian-Human immunodeficiency virus (SHIV) causes an increase in ACE2 expression in their cells as well. But, HIV and SHIV are not the only viruses that cause an interferon response and hence an increase in ACE2 expression. Humans infected with influenza A or B virus shows an increase in ACE2 in their cells. And it was not just the viral infected cells that showed an increase in expression of ACE2 but non-infected nearby cells showed this increase too.
ACE2 expression regulation in other diseases?
This brings me to yet another question. What about people with other non-viral co-morbidity? Can the interferon related increase of ACE2 explain some of the severity of the disease in these people? It is a good question to address. Well, at least a transcriptomic analysis of the lungs of patients with co-morbidities associated with covid19 show higher expression of ACE2 in their lungs (Bruna et al., 2020). But, it remains to be seen in which conditions this increase in ACE2 expression dependent on interferon at all. Maybe, we can have more discussion on it in the comment section.
Cell lines and mouse models
All that said and done, here is one more take-home message that comes out of this study by Zeigler et al., 2020. This is regarding the appropriate cell lines and animal model. For instance, in this paper, they were not able to find any increase in ACE2 expression in a bronchial epithelium - BEAS 2B cell line, on stimulation with interferon. However, they did see this happen in patient-derived cells. Well, cell lines are often transformed and change of the course of cell culture conditions. It happens many times that cell lines don't represent what happens in actual cells. And, patient-derived cells primary cells in culture are much more closer to their in vivo counterparts than cell lines. In fact, BEAS 2B cell line does not express ACE2, to begin with.
In fact, in this regard, I would like to mention this paper by Hoffmann et al., 2020 which can serve as a good guide for which cell line to use. A good human cell line as far as infection by SARS-CoV-2 is concerned would be Caco 2 (human epithelial colorectal adenocarcinoma cell line) or Calu 3. (lung cancer cell line). Vero E6, a cell line derived from African green monkey kidney epithelial cells, also serves as a good system. Moreover, it would be interesting to see if SARS-CoV-2 itself can induce an interferon response that increases ACE2 expression in these cell lines or in primary cells. And if that happens the inhibition of infection in, in vitro studies using cell lines that don't respond to interferon in a similar way may not produce an accurate picture. However, these cell lines would nevertheless be fine in screening for drugs that inhibit Spike-ACE2 binding, for instance.
Moreover, the authors found that mouse tracheal epithelial cells do not respond to interferon as human cells do. Interferon increased the expression of other interferon-stimulated genes in mouse cells but ACE2 should only a slight increase, Same was true for mouse treated with intranasal interferon spray. There was only a slight increase in the number of cells expression ACE2 and a slight elevation in the expression of ACE2 gene. So, this again makes one ask whether a mouse model infected with SARS-CoV-2 paint the exact picture for certain studies. For example, would an animal trial with interferon-gamma or interferon-alpha 2 in combating Covid19 a good model?
Anyhow, the regular mice itself is not a great model for SARS-CoV-2 by itself. There is a difference in the sequence of human and mouse ACE2 protein and hence Spike of SARS-CoV-2 doesn't bind as efficiently with mouse ACE2 as it does with Human ACE2 (see the link). Nevertheless, there is a transgenic mouse which comes to the rescue - K18hACE2 mice. This mouse is genetically modified to express the human version of ACE2 in airway epithelial cells (McCray et al., 2006). Since this mouse already has a high expression of human ACE2 in its airway, it is a good model to understand drugs that can inhibit viral binding and entry into the cells. It will even work fine for other downstream processes like viral replication inhibitors and viral protease inhibitors. However, when it comes to biologicals such as interferons or those that modulate the immune system to indirectly affect the virus, it must be better to evaluate that how much this mouse model mimics what happens in humans in response to a similar drug.
Final remarks
I think what authors found in this paper is definitely worth digging into further. One drawback of paper is that it mostly looks at RNA level of ACE2, but doesn't dwell much into questioning about protein levels of ACE2 in the cells of interest (Except one in vitro experiment). Moreover, I would love to know how interferon regulates ACE2. They mention that STAT1 might be regulating the expression of ACE2 in response to interferon. However, if that is the case then why this does not happen in mice? Is the promoter sequence for ACE2 different in humans vs mice? That is something someone needs to work on. Finally, I would like to know how ACE2 is regulated in Covid19 related comorbidities - Type 2 diabetes, hypertension, cardiovascular diseases, cancer, etc?
Anyhow, I guess that is it for today. We can continue the discussion in comments. Do share whatever comes to your mind.
About STEMSocial
But, before I go I would like to mention about the STEMSocial platform. Well, if you love reading and writing interesting science articles @steemstem or now STEMSocial is a community on hive that support authors and content creators in the STEM field. If you wish to support @steemstem do see the links below.
Or delegate to @steemstem.
25HP
50HP
100HP
200HP
250HP
500HP
1000HP
10000HP
100000HP
Also, if you have any questions regarding steemstem, do join the steemstem discord server.
You can DM me on discord, I have the same handle - @scienceblocks. Also if you are not a steem user, and reading this blog inspired you to start your science blog, find me on discord and let me know about you. I can try and help you navigate your way through steem.

Video by @gtg
References
Hwang et al., 2018. Single-cell RNA sequencing technologies and bioinformatics pipelines
Wu et al., 2020. Prolonged presence of SARS-CoV-2 viral RNA in faecal samples
Xiao et al., 2020. Evidence for Gastrointestinal Infection of SARS-CoV-2.
Wang et al., 2020. Detection of SARS-CoV-2 in Different Types of Clinical Specimens
Borden, Silverman and Sen. Chapter 52 - Interferons
Mice, hamsters, ferrets, monkeys. Which lab animals can help defeat the new coronavirus?
In that vain, as a layman in medicine and science and the scientific process and the scientific manual, It is my understanding that we have bacterial infections that are anti bacteria treatment resistant. This was, according to my understand, an over prescribing issue on the part of the Medical Professionals.
has there been any studies done about anti-viral treatment resistance? Is the reason for the increased difficulty treating viral infection due in part to the over use of anti-virus vaccines? I am not sure what the numbers are from the CDC, but every year they issue a new flu shot vaccine, and the majority of the years it is initially the wrong vaccine.
It seems to me that since the introduction of the flu shot, (vaccine), that the percentage of people that contract the flu has an increased death rate compared to 40 years ago. (I have not looked or checked or even know where to find that statistic).
Here's a link that goes into that a bit
https://respectfulinsolence.com/2020/03/31/coronavirus-viral-interference/
I looked through the link, while it was interesting it really did not answer or provide an answer of increased viral immunity to the flu vaccines, in the same respect as bacterial infections become more antibiotic resistant.
I do not believe the flu vaccine can cause an increased risk of contracting covid19, I do think a study needs to be done to determine if Viral infections are beginning to act like bacterial infections and becoming more resistant to the vaccines used to control them. The link mentioned nothing like that.
(hopefully this is not a double reply).
There is some interesting links inside this article here, especially the one on reverse manipulation of the virus if that is what this woman scientist really done as it pertains to covid, the big question on everyone's mind is did she develop this virus. Some claim she did then became infected and died, in handling her body not knowing what they were dealing when handling her body it infected others and supposedly she was patient zero. China denies this but no one has seen her since November, China claims she is alive and well but hasn't produced her to claim that is the truth. If it's a lab grown virus the debate over influenza and any influences is rather void.
https://esteem.app/gems/@kushfreeman/the-alternative-covid-19-narrative
There are four different classifications of viruses, Alpha and Beta, if I remember right corona viruses are classified as beta. You google what's the difference between influenza virus and corona virus it will run it down for you. Here's another link about vaccine research into corona virus. Don't know if all this helps, I know it sure is confusing to people who have no medical knowledge....that's for sure.
https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3371787/
Here's another link on the woman scientist:
The mechanism of resistance to antibiotics (the anti bacterial) drugs and to antivirals drugs are quite different. For instance a bacteria may acquire an enzyme capable of degrading the antibiotic. The viruses have a limited genome and I am not aware of them acquiring enzymes to degrade the antiviral drugs. Though, viruses mutate a lot. The antiviral drugs are designed to bind to some key molecule of the virus. However, it may so happen that a mutation in that protein molecule of the virus which happens, such that - its function remain intact, but the structure changes just a tat bit and decreases efficiency of antiviral drug to bind to it. In such a case this resistant strain will be selected for. Though anti viral drugs are not prescribed as much as antibiotics and this problem is not as big. But it might be of concern to patient being treated for chronic viral infections - HIV, Hepatitis, HSV, HPV etc. Strasfeld and Chou in this review describe some of the antiviral drug resistance mechanisms. You can follow up the references for further studies and let me know if you have any further questions on this. I will answer about the flu vaccines in next comment.
Here's what I been looking at some today. I guess from what I am gathering...am I am not in the medical field, is that interferon and hydroxychloroquine along with the antibiotic are used to break the cellar membrane to allow zine to access the cell. They act as an ionophore. Zinc inside cells stops replications of the virus but you have to have an ionophore to get zinc to be inter-cellar.
I did some looking up what ionophore meant,
ionophore [i´on-o-for″]
any molecule, as of a drug, that increases the permeability of cell membranes to a specific ion.
Miller-Keane Encyclopedia and Dictionary of Medicine, Nursing, and Allied Health, Seventh Edition. © 2003 by Saunders, an imprint of Elsevier, Inc. All rights reserved.
i·on·o·phore (ī-on'ō-fōr),
A compound or substance that forms a complex with an ion and transports it across a membrane.
[ion + G. phore, a bearer]
Farlex Partner Medical Dictionary © Farlex 2012
ionophore (ī-ŏn′ə-fôr)
n.
A compound that facilitates the transport of ions across a cell membrane, either by binding with the ion or by creating a channel through the membrane.
I looked at other articles on ionophores, I ran across words like quercertin and epigallocatechin which are used as natural ways to enhance introduction into a cell. There is some zinc inside cells naturally just not enough to ward off the replicating virus, we gain that zinc through foods we eat and we also naturally consume quercertin through certain foods we eat, quercertin also comes in a tablet form but they don't recommend taking more than 500 to 1000 mg a day, so you could increase your zinc intake and more of it would get into your cells by taking it or eating more foods that have it. Epigallocatechin on the other hand they do not recommend you take any amount in a tablet or power form without a doctors permission but it is found in abundance in green tea.
The clinical trials are being conducted for these. We have to wait and see the results. I also would not recommend that anyone takes any drug mentioned here without doctors advice. Hydroxychloroquine for instance have side effects including cardiac toxicity. Even in trials patients showing ecg distortions are taken off that drug. Moreover, the results of trials dont look very promising as of now.
O specifically dont want to do a post about these drugs because these desparate times and any false hope or wrong half information is not a great idea. I think lets wait for trials and pop anything only if your doctors tells you to.
However , if you want to have some green tea, sure I will make myself some too. Though I wont OD thinking of it some magic potion. More like who minds couple cups of green tea in a day.
I'll take the trial of a thousand patients in French with a 91 percent cure rate over anything Dr Fauci is peddling, if I am offered a chance to show improvement in four less days over being cured I am going to pick being cured. There another study gearing up in Australia right now, they to are using chloroquine with HIV drugs that show a one hundred percent cure rate. They are actually even calling it the cure all to the corona virus. They are sounding so confident in their finding that they are not even worried about a second wave.
I am like you though, I am not much into taking something that isn't prescribed and I think I would also opt for a good strong cup of green tea.
Well, the reason for needing a new flu vaccine every year is because of the existence of multiple strains. You don't know which strains are going to hit this year. Based on data on ongoing infections a flu shot for each year is designed. And it's not just the existence of multiple strains, all these strains are also pretty dynamic. They keep mutating and undergo antigenic shift (a mutation that won't allows antibody that recognises previous strain to bind to a new one). So a vaccine that worked last year for a particular strain may turn obsolete if that strain has undergone an antigenic shift.
The idea to avoid or at least minimize such drug resistance or antigenic shift is to design drugs or vaccines against highly conserved viral parts. These are parts that need to interact with host proteins for example. Hence, any changes in the structure via mutation in viral genome will severely distort the functioning of virus itself. At least that is what the aim usually is when picking a drug target or a subunit for vaccine. It's challenging but a sensible approach.
Finally, I am not aware of data on increase in flu deaths correlated to flu shots. Sounds doubtful. Do share a link for that data, I can have a look into it.
Also, thanks for reading my post and asking such interesting questions. Do leave them here if you have more.
A few weeks ago there was a paper published out of a hospital in China that said people with an active viral infection that getting the covid virus could accelerate that virus. There are viruses that if left unchecked can lead to encephalitis in the brain, what they suspected of one man who died whom they found the virus in his spinal fluids is that he had a primary virus infection and that covid possibly accelerated that infection. Since then I've read other articles of concern of the potential of this virus to accelerate viruses people get from mosquitoes and with the approach of summer coming on this issue was of concern to them.
Interesting, can you share the link or the author name for that paper. I would like to read it.
https://medium.com/microbial-instincts/neurology-and-covid-19-everything-researchers-know-so-far-f7e0607e2071
https://medium.com/microbial-instincts/first-case-of-covid-19-encephalitis-7b305fcfe9a8
Thanks for your contribution to the STEMsocial community. Feel free to join us on discord to get to know the rest of us!
Please consider supporting our funding proposal, approving our witness (@stem.witness) or delegating to the @steemstem account (for some ROI).
Thanks for including @stemsocial as a beneficiary, which gives you stronger support. Using the STEMsocial app could yield even more supporti next time.
Very nice journal club!
All of this is a bit weird. So, let me recap to see whether I got it right. If the COVID-19 virus uses the anti-viral response of the system to propagate, it is basically unstoppable, isn’t it? Therefore, how to get a treatment?
On the other hand, I also conclude from reading your article that anyone with an existing pathology may be more easily infected. Is that correct? Is there again anything we can do here?
This virus may be more violent than expected, no?
Well yes, if these results are to be believed then that is what it would imply. But of course there is a catch. Not all interferons raise ACE2 levels. Even in this study interferon a2 was much more potent than interferon gamma. So its not all lost. Moreover, the virus has a protein similar to protein found in SARS1. The orf9b of virus is a potential interferon antagonist. It remains to be seen if other than inhibiting antiviral response it also can act to limit severity in some cases. It may be a dynamical system, and eventual result may depend on multiple variables interacting. It needs to studied further.
And yes it seems that people with preexisting pathology have higher risk of contracting the disease and becoming severely ill. The higher ACE2 might be a good plausible explanation.
I think the best treatment approach would be to find small molecules that inhibit ACE2 and Spike binding. This is direction we are taking for now. Nonetheless, there can be other targets esp where protein of virus interacts with host proteins. I can do a post on that later.
Thanks for the clarifications (and raising the fine prints ;) )/
Cheers!
It is a very violent virus and easy to spread, perhaps preventive measures will help a little but not completely, sometimes it is inevitable not to get infected, now how close are we to being able to get a vaccine soon? I was reading that a group of scientists at Oxford University are very close to finding a vaccine, I ask you because I know you are in Europe and you are working with this. Unfortunately in our country we can't do much even though we have excellent professionals in the area of virology
That's a good question but also a difficult one to answer. I can't tell you when, but I can tell you that there is a lot going on for it. The trials are being conducted for something as novel as RNA vaccines. There are a few groups who have developed monoclonal antibodies against the virus and some more who are still in process. There are others who have claimed some success with inactivated virus in rats and macaques. But that's the thing most of it is in early development or early human trial. Even the Oxford one was tested to work in macaques. They are running phase 1 trials now. You can read this to see current state of some efforts.
Now the phase 1 trial will mostly test the safety and immunogenic potential (ability to mount immune response and produce antibodies) in small group. We have to wait at least until phase 3 to be sure if vaccine is both safe and effective against the virus. Some of these efforts would leas nowhere. Those that do may take at least a year (that is most hopeful scenario) to two (more realistic though still bit wishful scenario). If everything works out just fine, the approvals are accelerated then 1-2 years is wishful but doable. That is what I think is the scene.
Hello thank you for responding, I am aware that some tests are already being conducted and I am also aware that it takes a long time to find the right antidote, there are many tests, many phases, but I read in an article that they are already testing on people ... this is true ... I honestly do not think so and even more so for what you just told me, that maybe in a year we can have the vaccine.
Thank you for your answer and I apologize that I respond so late, my work is currently very demanding and I have little free time, which is why I enter very little stemsocial
Quite the same here. I hope to write and read posts every evening, but by the time I finish work I am always too late and tired to anything useful.
Yes, a year is wishful thinking for sure. But deep down I do hope that I am wrong and wishful thinking turns into reality.
@scienceblocks : I have a very fundamental question: Sorry if it is off-topic. How do virus acquire the lipids for its outer membrane? I know that all viruses have no outer covering. But sars-covid19 has one. Lipid synthesis or self assembly has nothing to do with genetic code from RNA/DNA(but the viral proteins are encoded in their RNA/DNA). Am I right? Can we target the virus while assembling?
Well there are lot of membrane viruses, aka enveloped viruses. HIV, flu, hepatitis, herpes. The virus take lipids from host membrane. For instance in case of corona virus the proteins - E, M and spike gets into ER. They are transported to the plasma membrane by ER - Golgi route. Somewhere in between the bind to the N protein along with the viral genome and gets wrapped around it.
In fact if you just transfected the cells with spike protein or infected it with a corona virus you will see the spike being expressed in plasma membrane and it causes the cells to fuse and form syncytia. The lipids these proteins interact with are what it tries to take along.
I dont think anyone has put much thought into it but yeah its worth a shot trying to target region on viral proteins that helps it interact with lipids. Though I am not sure if this can be done specifically and how conserved this target site would be. Alternatively, interaction between E, M and N, can be targeted. If a bioinfo study can point out existing drugs that targets anything that may disrupt virus assembly thd drug can be tried in lab for sure.