"Citizen Science" Project Activity Report 2 - Simulating top quark production at CERN’s Large Hadron Collider
"Citizen Science" Project Activity Report 2 - Simulating top quark production at CERN’s Large Hadron Collider
By Enio...
This article consists of a report of activities in the context of the participatory research project Citizen science particle physics project on Hive led by @lemouth. In it we will see the realization of the second set of activities of the project as described in this article and deals with the simulation of top quark production at CERN's Large Hadron Collider.
Task 1: Top-antitop production at the LHC
I went to the directory where I have the MG5_aMC installation and launched the software. Then I entered the command generate p p p > t t~ which, as we can see in the instructions, serves to define the production of a top-antitop pair in proton-proton collisions.
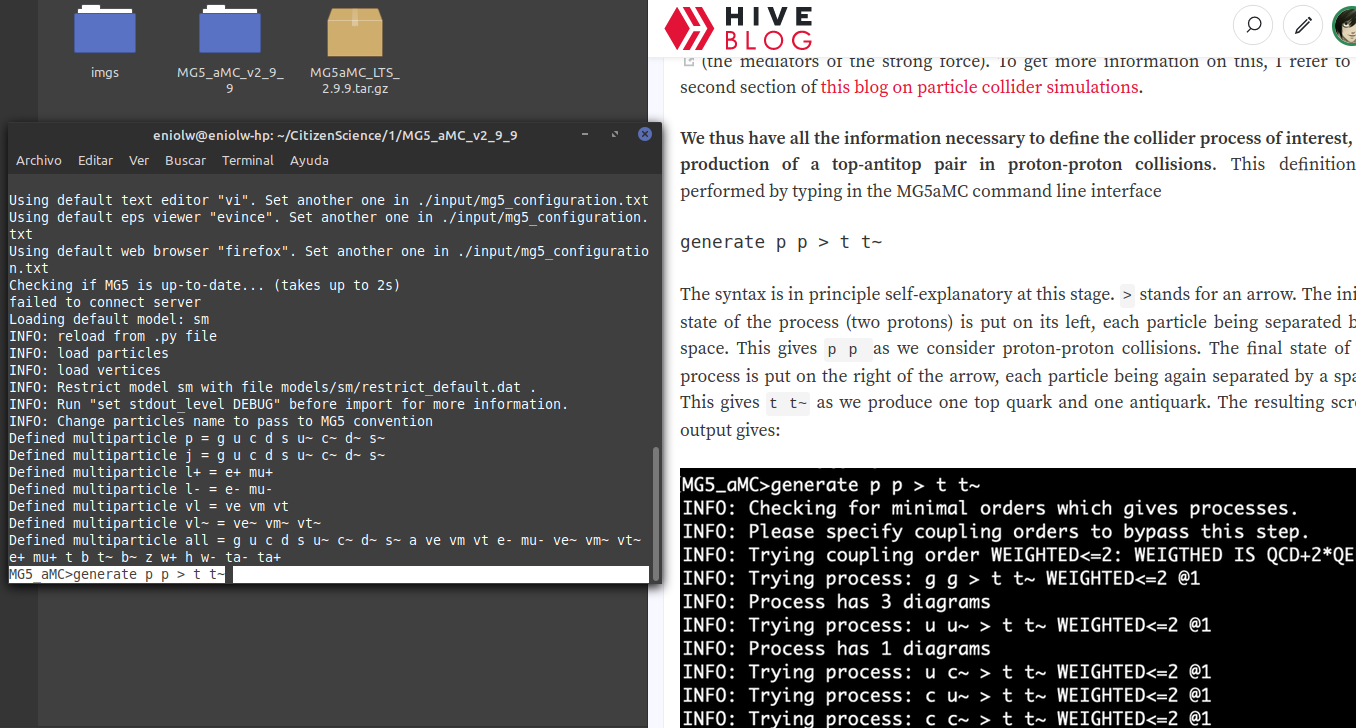
⬆️ Image 1: Entering the first command according to the instructions.
That produced the following output:
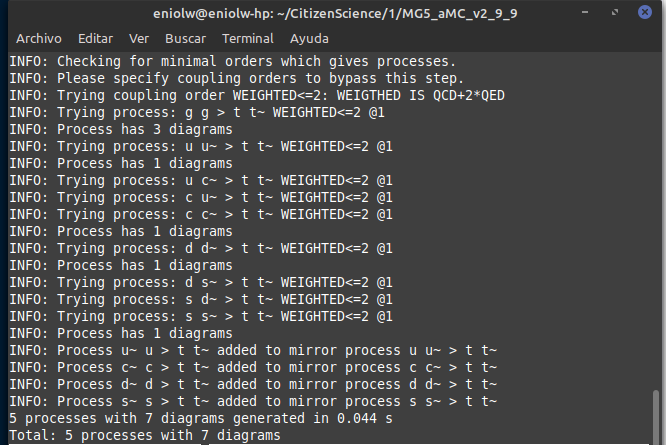
⬆️ Image 2: Everything is normal so far.
I then proceeded to generate the Feynman diagrams with display diagrams, which in turn produced the following output.
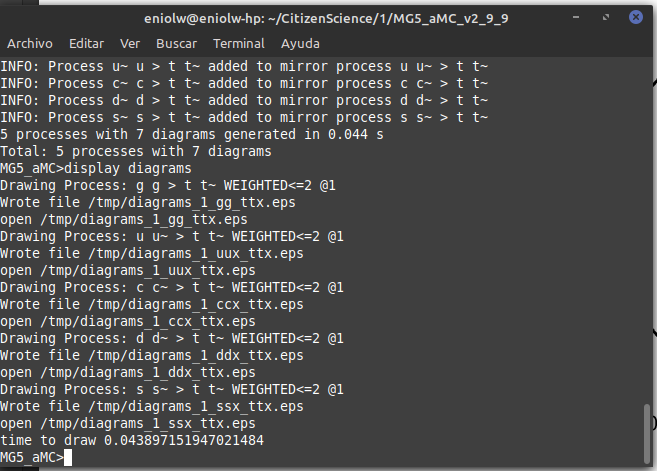
⬆️ Image 3: Second command output.
It says that several .esp files were generated in the /tmp directory. I took a look and indeed they are there. Below is a kind of thumbnail of the 5 files.
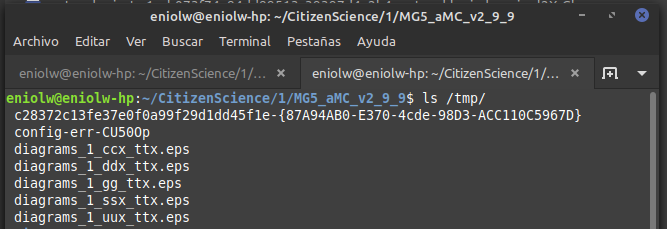
⬆️ Image 4: The /tmp directory showing the just generated files.
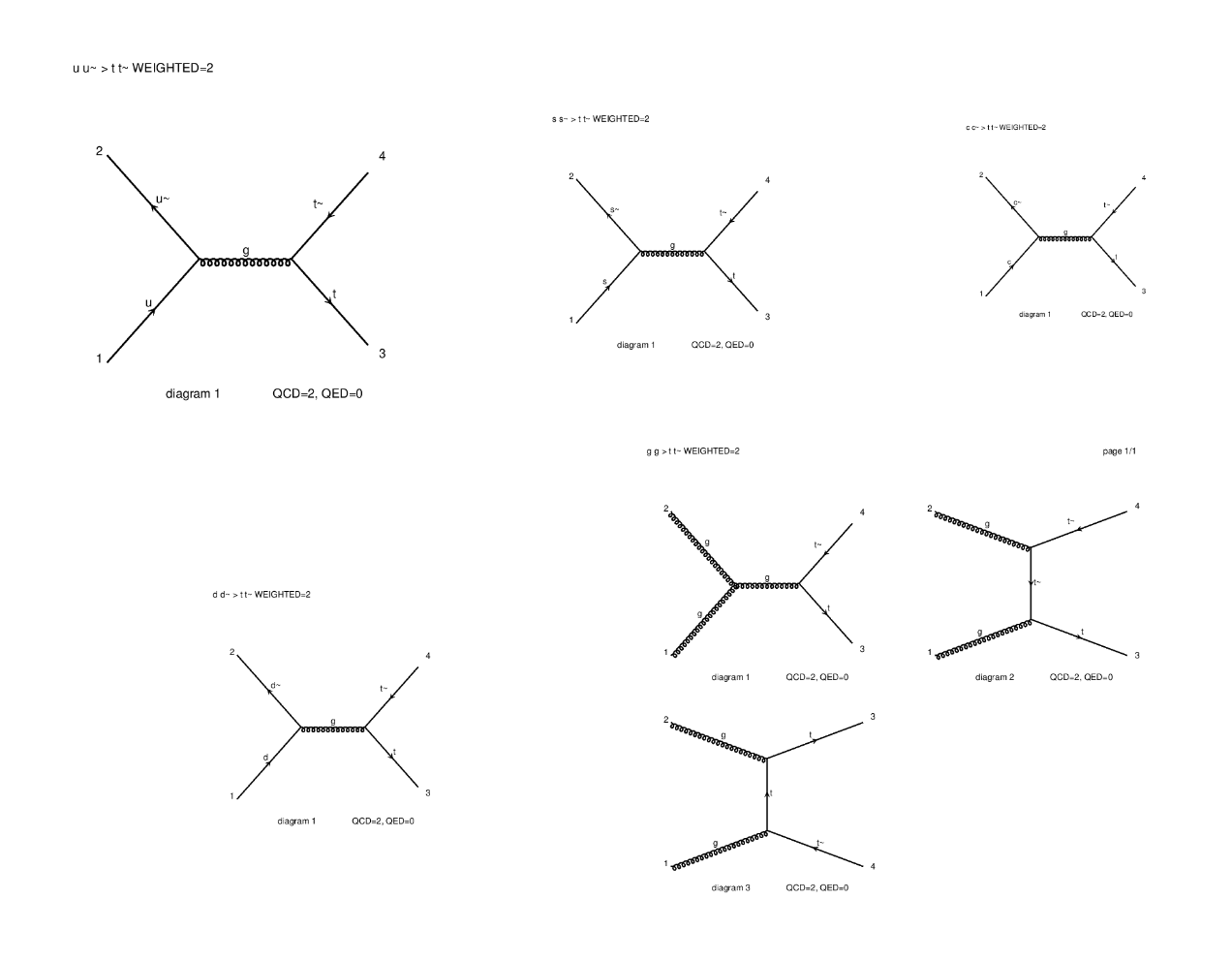
⬆️ Image 5: The five diagrams created (try opening them on another tab to zoom in).
I can't understand these images yet, but according to the instructions we don't need to interpret anything yet.
Task 2: A Fortran code for top-antitop production at the LHC
In order to extract an equation from these diagrams, I have entered the command output fortrancode1. This generates a directory named "fortrancode1" nested in the software directory. Using a new terminal tab I explored the file to verify that the files were generated.

⬆️ Image 6: Third command output.
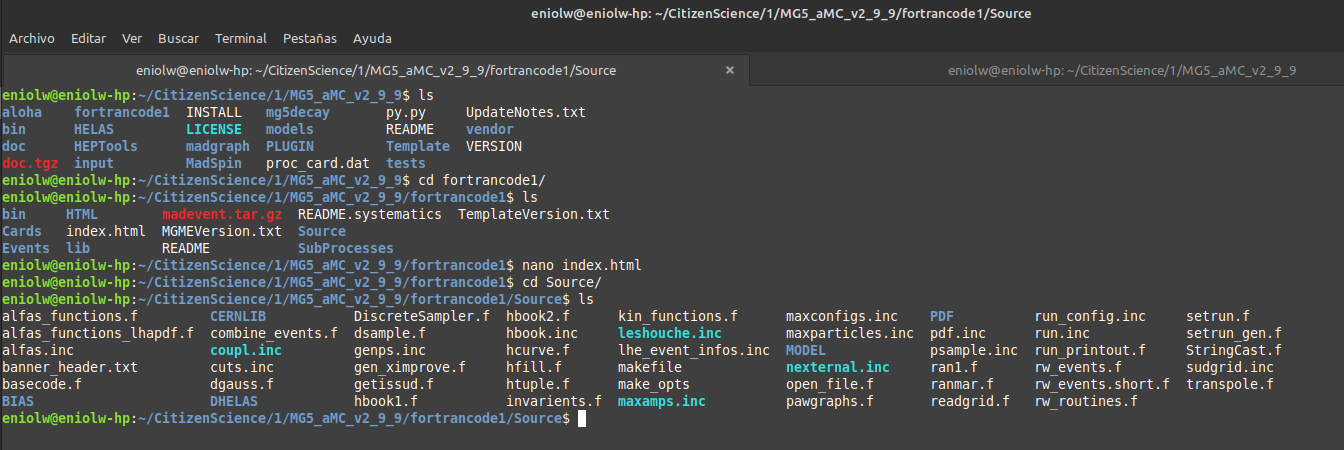
⬆️ Image 7: Checking the brand new directory "fortrancode1".
Out of curiosity I opened the dgauss.f file in the /source subfolder.
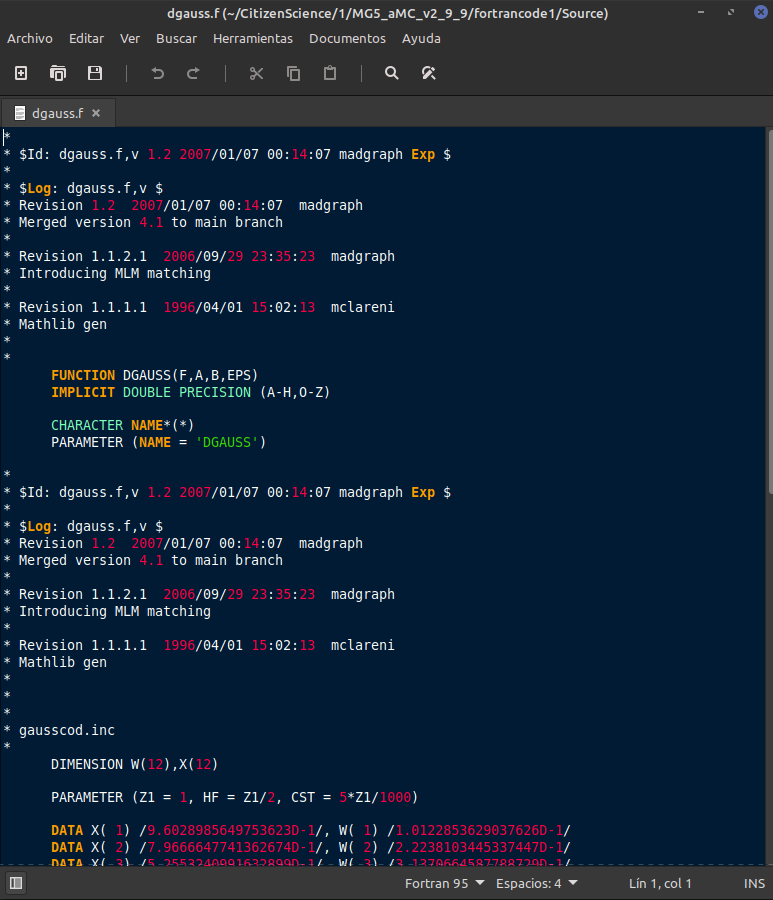
⬆️ Image 8: looking at one of the files out of curiosity.
Task 3: Computing the rate to top-antitop production at the LHC
I went back to MG5aMC to start the process of compiling the Fortran code with launch fortrancode1. This was a bit stressful and in the end I got a compilation error after two attempts. On the first attempt, following the instructions on the interactive menu that appears after using launch, I entered: 1, enter, 4, enter, but failed to edit the line 129 successfully via terminal. The edition process did not behave as expected and I guess it was a problem with Vim. I had to edit using xed, a GUI editor. After that it started running a process for several minutes, but in the end it failed and gave this output:
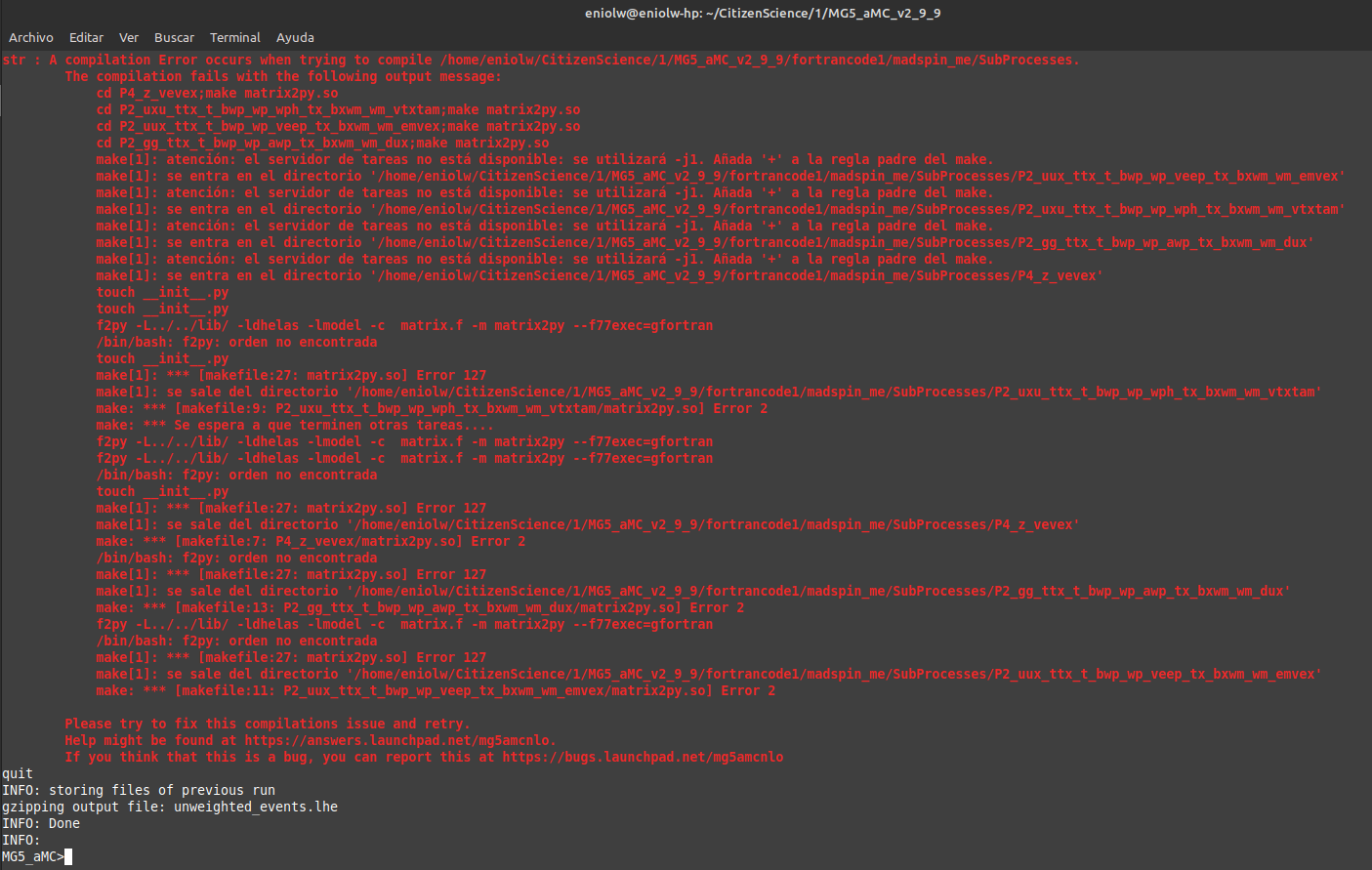
⬆️ Image 9: Something went wrong.
On my second try, I decided to set Nano as the shell text editor to see how it would behave, but in the process I seem to have mistakenly altered another parameter listed in the interactive menu. The truth is that this new compilation also failed. I guess this has something to do with the fact that I changed the line True = use_syst ! Enable systematics studies by False = use_syst ! Enable systematics studies and the tutorial warned that "the code should just crash because some packages are missing…". The point is, I'm not sure if this won't get me through task 4. @lemouth, I appreciate your guidance on this.
In any case, these processes produced an HTML file as output and was automatically opened by the web browser. I attach this view below:
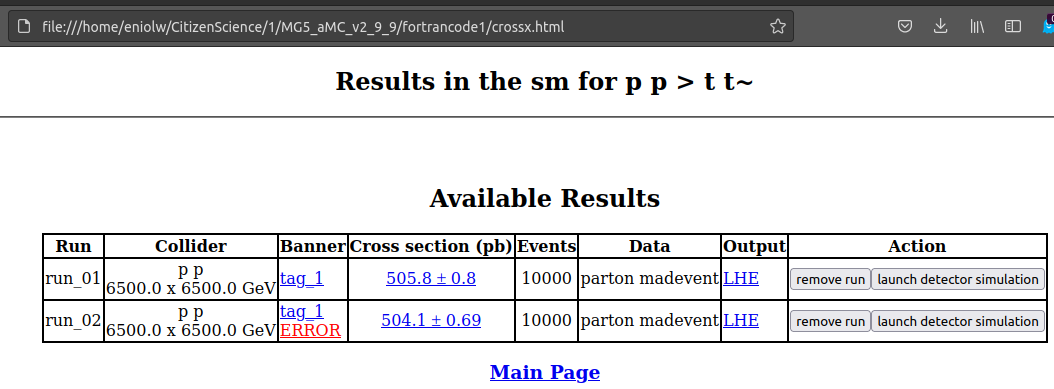
⬆️ Image 10: Simulation output?
This closely resembles the results of the tutorial, so it did work correctly, except for the part I mentioned.
Task 4: Checking out whether everything went fine
When I go to the fortrancode1/Events directory, I don't see the run_01_decayed_1 directory in it, but rather run_01.
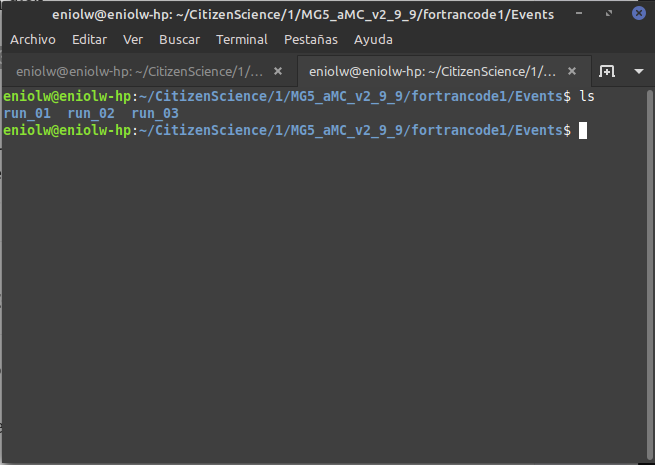
⬆️ Image 11: Actually, it's three directories. I guess I made three attempts after all.
When I go into them and list their contents, I don't see a tag_1_pythia8_events.hepmc.gz file, but this:
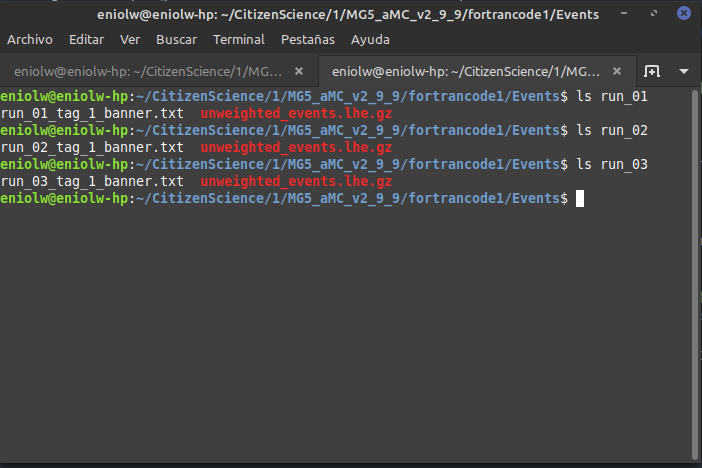
⬆️ Image 12: Something seems missing.
So I don't think I can say that this step is complete :( I hope this information is enough to diagnose the problem. I will be waiting for feedback from @lemouth to correct this and update the report. See you next time.
UPDATE #1 2022/04/05:
After @lemouth's feedback I was finally able to fix the problem by installing NumPy. (Can't believe I didn't have it installed on this machine.) The compilation process took a while and it overheated my computer, but in the end it seems to have worked out well.
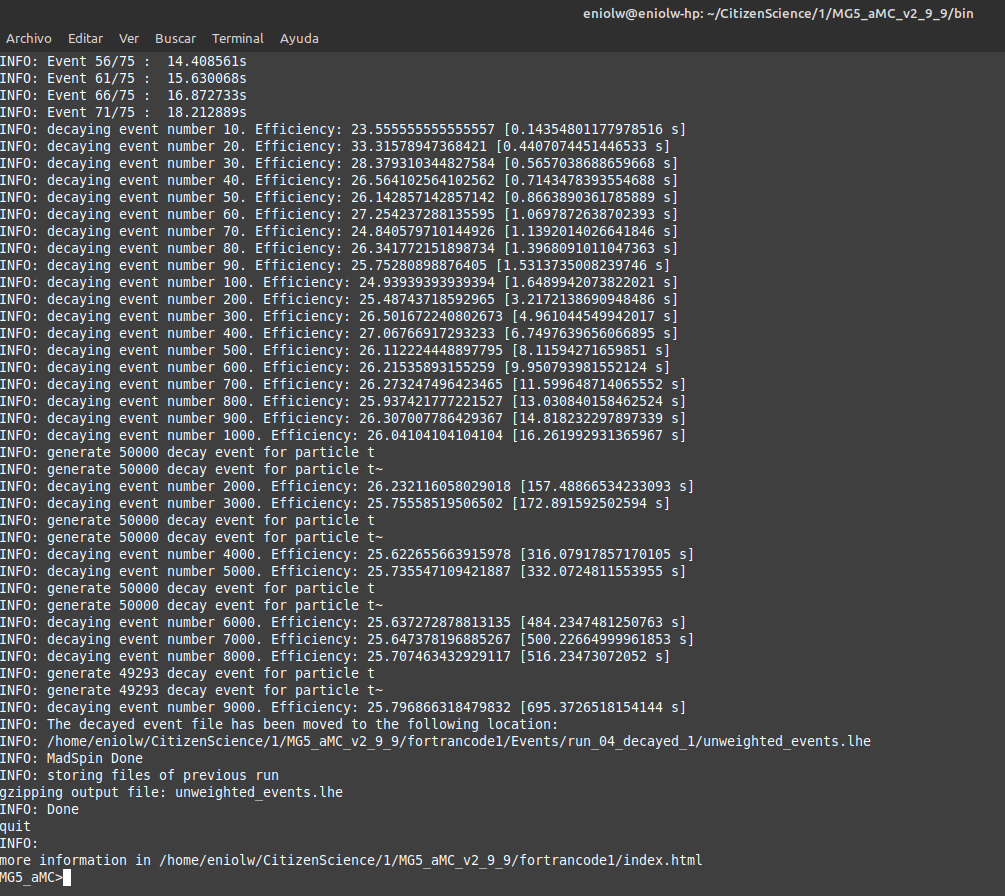
⬆️ Image 13: It's working.
However, I couldn't find the Pythia8 output in the log. So I guess it isn't complete yet. I'm pretty sure I enabled Pythia8 by pressing 1 as the instructions say, but I might be wrong, I don't know. What do you think, @lemouth? Should I repeat this? How?
UPDATE #2 2022/04/09:
After a few attempts and exchanges with @lemouth, I finally got the expected output. Apparently I messed up a bit my first MG5_aMC installation by changing the name of the root directory where I placed it (I had named it "CivilianScience" instead of "CitizenScience", so I changed it for the second project activity). I noticed it after @lemouth's suggestion to check the mg5_configuration.txt file. I fixed the root folder name in it, but still the program kept crashing. In the end I opted to do a brand new factory installation of MG5_aMC by repeating the procedure of the first activity. With this new installation everything behaved as expected. Below are some screenshots. Thanks, @lemouth, for your help.
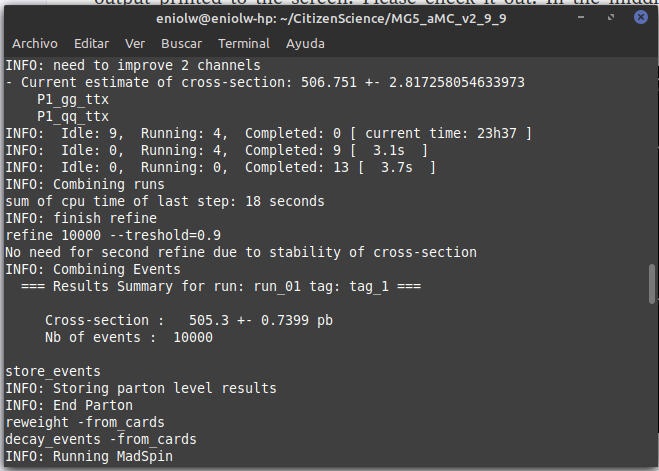
⬆️ Image 14: verification output
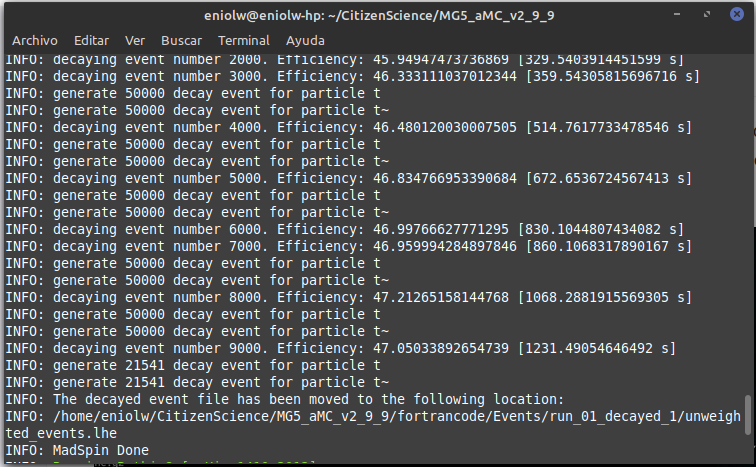
⬆️ Image 15: MadSpin done

⬆️ Image 16: Pythia8's output
Some References
If you are interested in more STEM (Science, Technology, Engineering and Mathematics) topics, check out the STEMSocial community, where you can find more quality content and also make your contributions. You can join the STEMSocial Discord server to participate even more in our community and check out the weekly distilled.
NOTES
- Unless otherwise stated, the images in this post are the author's.
Great Post!
!1UP
Thank you!
You have received a 1UP from @luizeba!
@stem-curator, @vyb-curator, @pob-curator, @neoxag-curatorAnd they will bring !PIZZA 🍕
Learn more about our delegation service to earn daily rewards. Join the family on Discord.
Thanks!
Thanks for this well-detailed report eniolw. I have a few comments, as you can expect, in particular due to the issue you found. I however also have comments relative to other parts of your report :)
First of all, you wrote:
I assumed this is a typo as you tried here to collide three protons (we do not have such experimental machines). In the screenshot, however, two protons are well there. Therefore, there is no problem here, I guess.
If you want a detailed explanation about these Feynman diagrams, here it is. I indeed already answered a related question of @mengene.
To provide some more insights directly targeting your question, those diagrams depict all ways allowed by the theory to connect the initial state (two constituents of the protons) to a top-antitop final state. The diagrams you showed in your report originate from quantum chromodynamics, the quantum theory of the strong interaction. Among all interactions included in this theory (that is part of the Standard Model), we have:
Those two interactions appear everywhere in the obtained diagrams, if you checked carefully. In practice, MG5aMC tries to construct all possible diagrams on the basis of the above two interactions. You can try manually and you should see that you would get the same graphs.
Note that any editor is fine, as long as you can edit the cards (and save the edited versions of them). The one that is chosen is at the end of the day only a matter of taste.
Now, let’s move on with your crash.
The problem here has nothing to do with your editing of the files (that was probably all right with both editors). It comes from the fact that the f2py package is not present on your system. It is a Fortran to Python interface generator, and it is part of NumPy. This package is required by MadSpin, that handles the decay of the unstable top quarks (so that we actually need it for the proposed tasks). As MadSpin was not used for the first episode of the project, we didn’t notice the issue so far.
I will edit my post to mention tat f2py needs to be present on the system. In the meantime, please install Numpy and retry again. Everything should be fixed. Please let me know.
Cheers!
I'm on it. I tried to compile again earlier, but surprisingly it took too much time, certainly over an hour. Is that normal? In comparison with your pictures, my instance output said it was counting like 50,000 events or something like that. I had to stop that process because I needed to work on other things. But now I'm going to let my laptop compute it all night. I will see in the morning if it's done. I hope so.
There are several weird things there.
Note that the run card can be directly verified at
There is thus no need to start the code to do so.
If all of the above is correct, maybe restarting from scratch, after having installed
f2pyis the thing to do. We will get there before episode 3, don't worry!Ok, the 50,000 probably had to do with some messages about the decay. I compiled way faster when I tried again. I updated the post. I got the MadSpin output but not Pythia8's. I'm pretty sure I enabled it. How can I repeat it? by doing the same with launch the_code?
From the screen output, it seems clear that Pythia has not run at all (therefore it is normal that you didn't file the output).
In principle, you could simply redo the launch command (
launch <folder name>) and verify that both Pythia and MadSpin are turned on.Note that at each new launch, the code will create a new
run_xxsub-folder. From therun_01,run_02,run_03, etc. list of folders, it is then sufficient to check the last one for the most recent run.Good luck!
Ok, I tried again. Please, take a look at this screenshot:
As you can see it says "shower = Not Available" for the option 1, which should be Pythia, according to your screenshot. But I did install NumPy. Am I missing something?
This means that
Pythia8has not been installed on the system, or has not been found. Can you please list the content of the HEPTools folder:You should have a
pythia8subfolder there, as well as aMG5aMC_PY8_interfacesubfolder (among other things).If this is the case, please have a look at
MG5_aMC_v2_9_9/input/mg5_configuration.txtand open this file.If not, please update the configuration file with the right paths (
< some path >is the absolute path of the folder in whichi MG5aMC has been installed).If none of this works, I am afraid that pythia8 will have to be reinstalled...
Cheers!
Thanks, it helped, but it kept crashing somehow.
I had to make a brand new installation and it worked. Check out my latest update in the post. :)
Finally! I am definitely happy to read this!
I have read the update of this blog. Indeed, changing folder names messes things up, in particular all paths and inter-relations between the packages installed through MG5aMC. In fact, a Pythia8 re-linking would have been sufficient here, but the full installation cleared that up.
See you thus on Tuesday for episode number #3 (which I should start writing soon) :)
Looking forward to it!
Thanks for your contribution to the STEMsocial community. Feel free to join us on discord to get to know the rest of us!
Please consider delegating to the @stemsocial account (85% of the curation rewards are returned).
You may also include @stemsocial as a beneficiary of the rewards of this post to get a stronger support.
Thank you!